Reprinted with permission. Biomedicines. 2025;13(9):2142.*
by Peter Manolakos, Yu-Bo Wang, Janice Withycombe, Luigi Boccuto, and Diana Ivankovic
Dr. Manolakos is with Healthcare Genetics and Genomics PhD Program, Clemson University in Clemson, South Carolina. Dr. Bo-Wang is with School of Mathematical and Statistical Sciences, Clemson University in Clemson, South Carolina. Drs. Withycombe, Boccuto, and Ivankovic are with Department of Healthcare Genetics and Genomics, Clemson University in Clemson,
South Carolina.
*Copyright: © 2025, the authors. Licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/). The article may have been modified to adhere to journal style.
FUNDING: No funding was received for this article.
DISCLOSURES: PM is an employee of and a stockholder in Novartis®. Novartis® had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results. Y-BW, JSW, LB, and DI declare no conflicts of interest.
ABSTRACT: Background/Objectives: KRAS mutations are among the most prevalent oncogenic drivers in non-small cell lung cancer (NSCLC), with their impact on survival influenced by comutations. SMARCA4 mutations are increasingly associated with poor prognosis and can be classified as class 1 or class 2 mutations. This study evaluates the prognostic implications of KRAS and SMARCA4 mutations, including their comutations and their impact on patients with NSCLC by utilizing real-world evidence. Methods: A retrospective analysis was conducted using the AACR GENIE Biopharma Collaborative (BPC) NSCLC 2.0 dataset. Patients with NSCLC with KRAS mutations, SMARCA4 mutations, or KRAS/SMARCA4 comutations were identified. Survival outcomes were assessed using univariate and multivariate Cox proportional hazards models, incorporating key clinical variables such as sex, race, smoking history, and stage. Results: Among 659 patients with NSCLC with KRAS or SMARCA4 mutations analyzed, KRAS mutations were the most prevalent (79%, n=518). SMARCA4 mutations were identified in 14% of cases (n=95) across two classes, 6% (n=41) with class 1 mutations and 8% (n=54) with class 2. Neither SMARCA4 class was associated with worse survival outcomes compared to KRAS-mutated patients (p=0.438 and 0.720). Patients harboring KRAS/SMARCA4 class 1 comutations (3%, n=18) had significantly worse overall survival compared to those with KRAS mutations alone (hazard ratio [HR]: 3.23, p<0.001). In contrast, KRAS/SMARCA4 class 2 comutations (4%, n=28) did not significantly impact survival compared to KRAS-mutated patients (HR: 1.34, p=0.205). Conclusion: KRAS/SMARCA4 class 1 comutations are associated with significantly worse overall survival compared to KRAS-mutated NSCLC. Our multivariate analysis demonstrates the critical need to incorporate routine next-generation sequencing (NGS) testing in managing patients with NSCLC at the time of metastatic diagnosis, with particular emphasis on identifying SMARCA4 mutation class as a potential prognostic biomarker in those with KRAS comutations. KEYWords: KRAS, SMARCA4, mutation, comutation, KRAS/SMARCA4, NSCLC, survival, prognostic, biomarker, AACR GENIE
Introduction
Lung cancer is the most prominent cause of cancer mortality globally among both men and women, with approximately 1.8 million deaths in 2023.1,2 In the United States (US), approximately 350 people die each day from lung cancer, causing more deaths in 2020 than prostate, breast, and pancreatic cancers combined.3 Non-small cell lung cancer (NSCLC) accounts for 84% of lung cancer cases, and the five-year survival rate was only 25% across all stages from 2013 to 2019.4,5 With the adoption of genomic testing and advances in targeted therapy treatment for patients with NSCLC, there has been some improvement compared to the standard of care in overall survival (OS) rates and progression-free survival.6,7 However, there needs to be a better understanding of how established NSCLC driver mutations, passenger mutations, and comutations impact OS outcomes.
In Western countries, mutations in the KRAS gene are the most identified driver mutation in nonsquamous NSCLC, presenting in approximately 30% of adenocarcinoma cases.8 KRAS mutations are associated with poor OS rates among patients diagnosed with NSCLC and have been determined to be a weak but valid prognostic biomarker.9 In relation to the current standard of care in oncology treatments, KRAS mutations in NSCLC have shown mixed results as predictive clinical biomarkers for survival outcomes with immune checkpoint inhibitor (ICI) treatments.10 More common KRAS comutations may impact therapy as retrospective analyses and critical reviews have posited that patients with NSCLC with KRAS/STK11 comutations are likely to exhibit primary resistance to ICI treatments.11 Questions arise when understanding the prognostic impact of other KRAS comutations with nondriver mutations in patients with NSCLC.
SMARCA4 mutations are found in approximately 8% to 12% of patients with NSCLC.12–14 SMARCA4-mutated patients tend to present with adenocarcinoma, a smoking history, and a low frequency of EGFR/SMARCA4 comutations.12,13,15 In addition, a comprehensive critical review encompassing 21 studies concluded that OS outcomes were worse for patients with NSCLC with SMARCA4 mutations than those with wildtype SMARCA4 alleles (tumors without SMARCA4 mutations).16
SMARCA4 mutations are categorized into two classes, as described by Schoenfeld et al, both of which are associated with epigenetic dysregulation but affect chromatin remodeling through distinct mechanisms.13,17 For example, class 1 mutations involve truncating mutations (ie, frameshift, nonsense), fusions, and homozygous deletions. SMARCA4 encodes brahma-related gene 1 (BRG1) protein, and class 1 mutations are often associated with BRG1 loss and loss of function.16,18 Class 1 mutations are mainly observed in NSCLC, and some cancer cases indicate that this loss of function leads to reduced chromatin accessibility and diminished remodeling activity.19 Class 2 mutations include missense mutations, which are suggested to induce gain-of-function or dominant-negative effects, possibly leading to deleterious cellular processes.
Two of the most extensive SMARCA4 mutation datasets categorized these classes and types of mutations in a similar percentage of patients. For example, in the analysis conducted by Schoenfeld et al, including 408 patients with SMARCA4-mutated NSCLC, 52% were class 1 and 48% were class 2.13 In this analysis, patients with SMARCA4 class 1 mutations had a worse prognosis and OS than patients with class 2 mutations or who were SMARCA4 wildtype (p<0.001). Moreover, the landmark analysis conducted by Dagogo-Jack et al of 3,188 SMARCA4-mutated patients yielded similar distributions, with 49% of SMARCA4-mutated patients being categorized as class 1 and 51% of patients as class 2.12
In the American Association for Cancer Research (AACR) Project Genomics Evidence Neoplasia Information Exchange (GENIE) Biopharma Collaborative (BPC), patients with SMARCA4-mutated NSCLC have an approximate 30% incidence of KRAS/SMARCA4 comutations.20 Moreover, KRAS/SMARCA4 comutated patients across multiple NSCLC studies were found to have inferior survival outcomes throughout various types of cancer treatment analysis, including chemotherapy, ICI, and targeted therapy.21–25 These inferior survival outcomes were also reinforced by the most extensive real-world evidence (RWE) retrospective NSCLC analysis of SWI/SWF mutations, including SMARCA4 variants. Investigators found that the patients with comutated KRAS/SMARCA4 NSCLC had worse survival versus with KRAS-mutated/SMARCA4-wildtype patients (hazard ratio [HR]: 1.882, p<0.00001).14 In contrast, one large analysis has demonstrated superior OS for patients with NSCLC with SMARCA4 mutations treated with ICI.13
A limited number of SMARCA4-mutated NSCLC survival analyses have been completed, directly comparing class 1, class 2, and SMARCA4-wildtype patients.13,19,21 As previously noted, there are several analyses comparing KRAS/SMARCA4-comutated to KRAS-mutated/SMARCA4-wildtype patients.21–25 To our knowledge, only one large multivariate analysis has comprehensively assessed KRAS/SMARCA4 class 1 and KRAS/SMARCA4 class 2 versus KRAS-mutated/SMARCA4-wildtype patients.13 Analyzing high-quality observational RWE phenotypic data alongside genomic mutations and comutations in NSCLC may better elucidate how KRAS, SMARCA4 (class 1 and 2), and KRAS/SMARCA4 comutations intersect and influence overall patient outcomes and survival, thereby informing future drug development. This study aims to retrospectively analyze the prognostic impact on survival outcomes of KRAS mutations and SMARCA4 (class 1 and 2) mutations or KRAS/SMARCA4 comutations in patients with NSCLC via the AACR GENIE BPC dataset.
Materials and methods
We utilized the AACR GENIE BPC NSCLC 2.0-public cohort, a publicly available dataset in cBioportal, to identify all patients with NSCLC with KRAS mutations, SMARCA4 mutations, or KRAS/SMARCA4 comutations.26,27 The NSCLC BPC cohort comprises 2,004 samples from 1,846 patients, randomly selected from samples in the GENIE 11.1-public release. The Dana–Farber Cancer Institute, Memorial Sloan Kettering Cancer Center, University Health Network, and Vanderbilt-Ingram Cancer Center contributed these samples. To be included in this analysis, patients were required to have at least two years of follow-up and a genomic sequencing report performed between 1 January 2014, and 31 December 2017. Clinical information was abstracted at each institution using the PRISSMM framework, deidentified, and provided to AACR for compilation.28 The dataset of KRAS and SMARCA4 mutations included all NSCLC histological subtypes and stages in the BPC cohort (ie, lung adenocarcinoma and lung squamous cell carcinoma). The study was reviewed by Clemson University (Institutional Review Board #2023-0636) and deemed exempt as it utilized a publicly available database containing de-identified patient data.
SMARCA4 class 1 and class 2 mutation categorizations were defined following the landmark analysis by Schoenfeld et al.13 SMARCA4-mutated patients with a frameshift deletion, frameshift insertion, nonsense mutation, fusion, splice site mutation, or splice region mutation were categorized as SMARCA4 class 1 mutations. Patients with missense mutations, in-frame insertions, or deletions were classified as SMARCA4 class 2 mutations. Tumors with concurrent SMARCA4 class 1 and SMARCA4 class 2 mutations were categorized as class 1. When multiple samples carrying the same mutation were attributed to the same patient ID, one sample was used for analysis, and the other samples were excluded. Similarly, patients with both KRAS and SMARCA4 mutations were merged under a single de-identified patient ID to avoid duplication in the analysis.
Smoking history was classified into three groups: never, current, or former user. Former users were combined [(quit <1 year), (quit >1 year), or (quit=unknown time)]. Disease stages were classified into stage IV versus stage I to III. Any patients that were stage I to III not otherwise specified (NOS) were classified as stage I to III. Patients were either categorized as White or non-White. Non-White patients included Black, Chinese, Other Asian, American Indian, or Other. Patients with “unknown” defined variables for categories in scope were excluded from the analysis to maintain the accuracy and interpretability of the results. A very low number of samples fell into the unknown category: race (n=18), stage (n=1), and smoking history (n=2).
Statistical methods. Statistical analyses were performed under the R software 4.4.1, and the statistical significance level was set at α equals 0.05. To assess the marginal association of each risk factor with survival status (the event is deceased), the t-test and Chi-squared test were first conducted separately for a continuous (ie, age and categorical [ie, sex]) factor. The results were presented along with other vital descriptive statistics (ie, sample means and frequencies). Then, the Cox proportional hazards model was fitted to present the final findings jointly for all these factors (ie, their joint associations with the deceased).
Results
After applying inclusion and exclusion criteria, 659 patients were retained in the dataset.
Clinical and genomic characteristics. Table 1 describes the clinical and genomic patient characteristics of the 659 patients with NSCLC. The mean age for next-generation sequencing (NGS) was 67.4 years. There were more female patients (n=397, 60%) than male patients (n=262, 40%). The most prevalent race was White (n=600, 91%) versus non-White/Other (n=59, 9%), with Black (n=32) and Chinese/Other Asian (n=16) as the most common non-White patients. Forty-one percent of patients (n=267) had stage IV NSCLC versus 59% of patients (n=392) having early or locally advanced disease (stage I–III). For early or locally advanced disease, stage I was the most common (n=192), followed by stage II (n=74), stage III (n=120), and stage I to III NOS (n=6). The vast majority of patients had a smoking history: former (n=496, 75%) and current (n=113, 17%) versus never having a smoking history (n=50, 8%).
Regarding the KRAS and SMACAR4 mutation status for all 659 patients with NSCLC, KRAS (n=518, 79%) was the most common (Table 1). There were more SMARCA4 class 2 mutations (n=54, 8%) than SMARCA4 class 1 mutations (n=41, 6%). Similarly, slightly more KRAS/SMARCA4 class 2 comutations (n=28, 4%) than KRAS/SMARCA4 class 1 comutations (n=18, 3%) were reported. Seven SMARCA4 class 1 patients had more than one type of SMARCA4 mutation and were classified as class 1 due to one of the mutations being a frameshift or nonsense mutation. One KRAS/SMARCA4 comutated patient had more than one type of SMARCA4 mutation and was classified as KRAS/SMARCA4 class 1 because they had a SMARCA4 frameshift deletion and a SMARCA4 missense mutation.
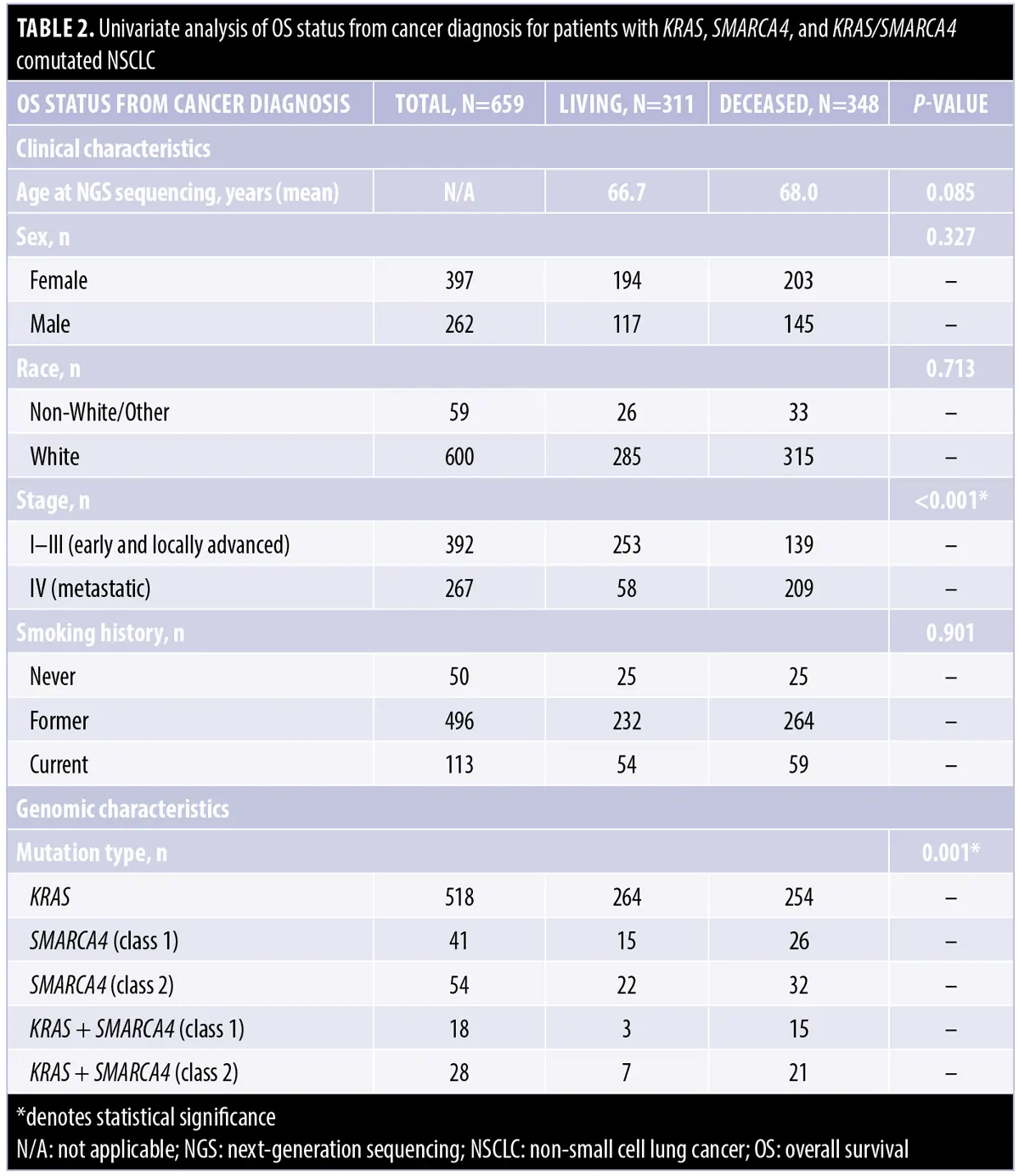
Univariate OS analysis. Table 2 provides the clinical and genomic univariate analysis of OS status (living vs. deceased) from cancer diagnosis. Factors of interest were evaluated for their marginal associations with survival status. Significant associations with the risk of death were found under α equals 0.05 in the factors “Stage” (p<0.001) and “Mutation Type” (p=0.001). While the mean ages of NGS sequencing were 66.7 and 68.0 years for living and deceased patients, respectively, the difference in the mean ages is insignificant (p=0.085). Additionally, no significant OS differences were found between sexes (female vs. male, p=0.327) or racial groups (White vs. non-White/Other, p=0.713) or even among patients with varied smoking histories (never vs. former vs. current smokers, p=0.901).
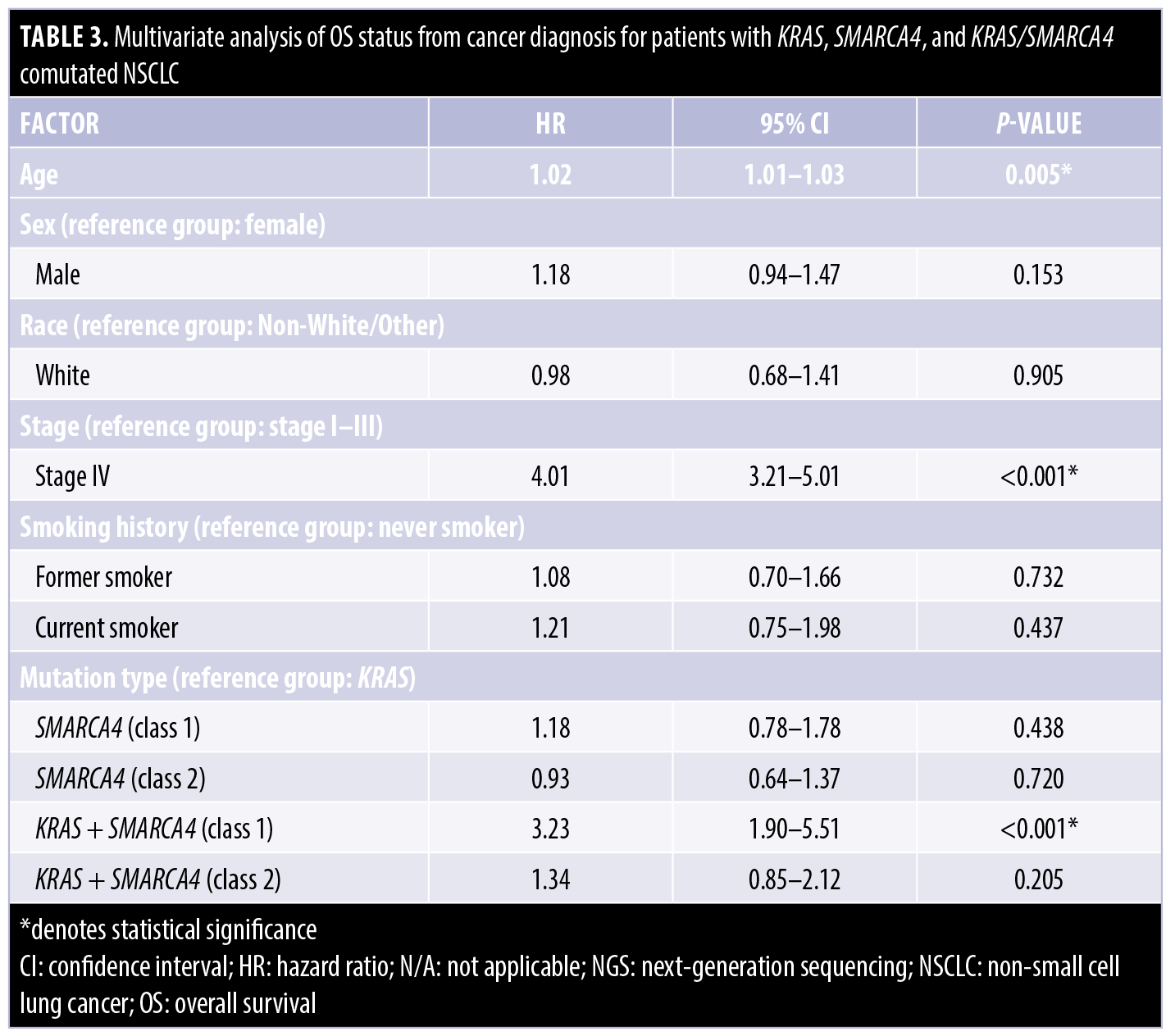
Multivariate analysis. Table 3 presents the results of the Cox regression model for patients with KRAS, SMARCA4, or KRAS/SMARCA4 comutated NSCLC. In this joint modeling, three factors were identified as important predictors accounting for a significantly increased risk of death. Specifically, “Stage IV” patients had an approximately four-fold risk of death than the earlier stages (HR: 4.01; 95% confidence interval [CI]: 3.21–5.01; p<0.001). KRAS/SMARCA4 (class 1) comutated patients had a 3.2-fold risk of death compared to KRAS-mutated patients (HR: 3.23; 95% CI: 1.90–5.51; p<0.001). The age at sequencing was also associated with an increased risk of death (HR: 1.02; 95% CI: 1.01–1.03; p=0.005).
Sex and race were not significantly associated with the risk of death [(male vs. female: HR: 1.18; 95% CI: 0.941–1.47; p=0.153) and (White vs. non-White/Other: HR: 0.978; 95% CI: 0.676–1.41; p=0.905)]. Similarly, smoking history showed no significant association [(former smoker vs. never smoker: HR: 1.08; 95% CI: 0.702–1.66; p=0.732) and (current smoker vs. never smoker: HR: 1.21; 95% CI: 0.745–1.98; p=0.437)].
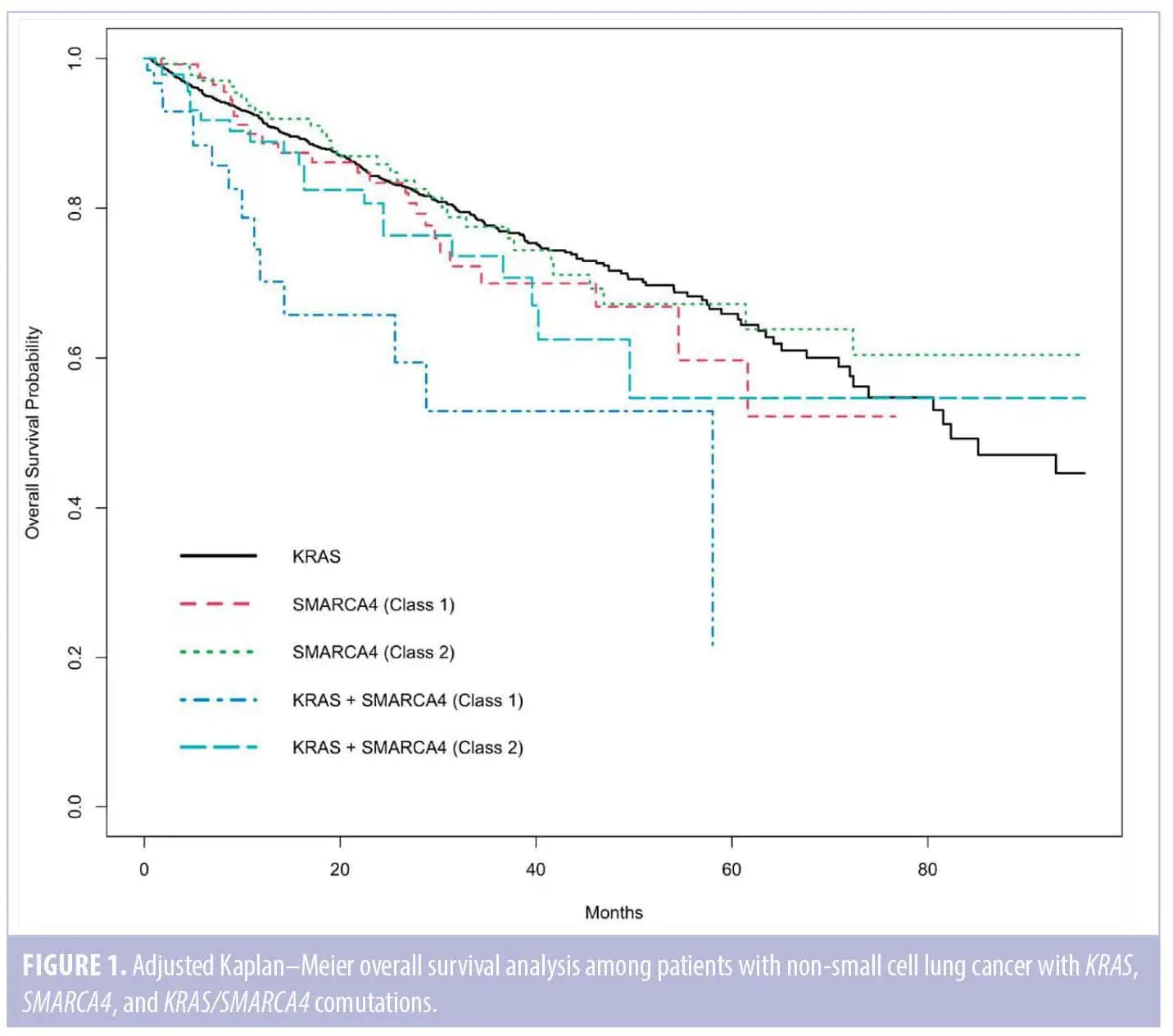
The other comparisons among the genomic mutation types were not associated with a significantly increased risk of death. Neither class 1 nor class 2 SMARCA4 mutations had significantly worse OS compared to KRAS mutations: [SMARCA4 (class 1) vs. KRAS (HR: 1.18; 95% CI: 0.779–1.78; p=0.438)] and [SMARCA4 (class 2) vs. KRAS (HR: 0.932; 95% CI: 0.635–1.37; p=0.720)]. The comparison between patients with KRAS/SMARCA4 (class 2) comutations versus KRAS mutation was not significant (HR: 1.34; 95% CI: 0.851–2.12; p=0.205). To summarize these findings visually, the adjusted Kaplan–Meier curves (adjusted for other risk factors) for these five mutation types are shown in Figure 1.
Discussion
Our retrospective RWE analysis provides a comprehensive view of the prognostic impact of KRAS and SMARCA4 mutations, including their comutations, on OS in patients with NSCLC. Utilizing the AACR GENIE BPC dataset, we identified distinct differences in genomic profiles influencing prognostic outcomes across a multivariate analysis. Notably, our findings highlight how these mutations and their comutations are significantly associated with increased patient risk, offering potential insights into how they may influence OS outcomes in NSCLC.
The results of the Cox regression multivariate analysis demonstrated that KRAS/SMARCA4 class 1 comutations are associated with worse OS compared to KRAS mutations for patients with NSCLC (HR: 3.23; 95% CI: 1.90–5.51; p<0.001). In contrast, KRAS/SMARCA4 class 2 comutations did not appear to confer a significantly worse OS than KRAS mutations (HR: 1.34; 95% CI: 0.851–2.12; p=0.205). In addition, SMARCA4 class 1 and class 2 mutations alone were not associated with worse OS when compared to patients with KRAS-mutated NSCLC.
Our findings support the possible need to determine the SMARCA4 mutation class as a prognostic factor when a patient carries a KRAS/SMARCA4 comutation. Moreover, a recent meta-analysis outside of the comutation scope posited that SMARCA4-mutated class 1 was associated with worse OS for patients with NSCLC (HR: 1.63; 95% CI: 1.44–1.85; p<0.00001) in contrast to SMARCA4-mutated class 2 where no OS association was observed (HR: 1.34; 95% CI: 0.87–2.06; p=0.18).29 This meta-analysis by Wankhede et al differs from our OS analysis because it did not directly compare KRAS/SMARCA4 comutated patients across both SMARCA4 mutation classes.
Although multiple NSCLC studies demonstrate worse OS in patients with KRAS/SMARCA4 comutations compared to KRAS-mutated patients, most do not analyze the impact of SMARCA4 class 1 versus class 2 on OS within this comutated population.22,24,25 For example, in the most extensive RWE analysis to date, Herzberg underscored the unfavorable prognostic effect of KRAS/SMARCA4 comutations compared to KRAS mutations (HR: 1.882, p<0.00001), and the SMARCA4 mutation class effect was not directly noted.14 They took a different approach and categorized SWI/SWF mutations as likely pathogenic or pathogenic (LP/P) per OncoKB, a precision oncology mutation database that provides the clinical significance of cancer-related mutations. Fifty-two percent of the SMARCA4 mutations used for their analysis were deemed LP/P and most likely included some missense mutations, as OncoKB does categorize specific types of missense mutations as LP/P. In contrast, our analysis categorized all missense mutations as class 2 regardless of LP/P status following the methodology applied by Schoenfeld et al.13
Similar to our results, Schoenfeld et al reported that patients with NSCLC with KRAS/SMARCA4 class 1 comutations were significantly associated with poorer survival (HR: 1.59; 95% CI: 1.04–2.41; p<0.001) compared to KRAS-mutated/SMARCA4-wildtype tumors.13 Additionally, their NSCLC analysis showed that KRAS/SMARCA4 class 2 comutations were associated with worse OS (HR: 2.75; 95% CI: 1.84–4.11; p<0.001) compared to KRAS/SMARCA4-wildtype tumors. These results remained prognostic after accounting for variables such as age, sex, histology, smoking status, TMB, and the presence of STK11 or Kelch-like ECH-associated protein 1 (KEAP1) comutations.
Biological insights. These survival findings reinforce the hypothesis that the loss-of-function nature of SMARCA4 class 1 mutations may lead to diminished chromatin accessibility and remodeling, impairing gene regulation and potentially contributing to tumor progression.30 As demonstrated in the Dagogo-Jack et al, Alessi et al, and Schoenfeld et al studies, most of the class 1 SMARCA4 truncating mutations result in a loss of BRG1 protein function.12,13,21 For example, in these three studies, 84% (26 of 31), 100% (11 of 11), and 81% (50 of 62) NSCLC samples with truncating SMARCA4 mutations lacked BRG1 immunohistochemistry (IHC) protein expression, respectively. These variants leading to BRG1 protein loss comutated with KRAS may lead to cancer treatment resistance by creating an immunosuppressive tumor microenvironment and impairing the effectiveness of deoxyribonucleic acid (DNA) repair mechanisms. Liu et al found that KRAS/SMARCA4 comutated patients had significantly lower activated cluster of differentiation (CD4+) memory T cells (p=0.0035) and (CD8+) T-cells proportions (p=0.015) than patients with KRAS-mutated/SMARCA4 wildtype NSCLC.25
Emerging therapeutic strategies for SMARCA4-mutated NSCLC. Currently, there are several therapeutic approaches targeting SMARCA4 mutations. SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily A, member 2 (SMARCA2) selective protein degraders, such as PRT3789, offer a unique approach by exploiting synthetic lethality in SMARCA4-deficient tumors with over 1,000-fold selectivity for SMARCA4-mutated cancer cells compared to wildtype cells.31 Two patients with NSCLC treated with PRT3789 had a confirmed partial response.32 In treatment combination approaches, ataxia–telangiectasia and Rad3-related protein (ATR) inhibitors are being studied in combination with ICI treatment in previously treated patients with NSCLC.33 Tuvusertib (M1774) is a selective ATR inhibitor with antitumor activity as monotherapy in preclinical models with DNA damage repair pathway gene mutations. It is being studied with cemiplimab in a cohort of patients with NSCLC with SMARCA4 mutations.33,34
Clinical implications. Our multivariate analysis underscores the critical need to incorporate routine NGS testing in managing patients with NSCLC at the time of metastatic diagnosis, with particular emphasis on determining SMARCA4 mutation class in those with KRAS comutations. We demonstrated that patients with NSCLC with KRAS/SMARCA4 class 1 comutations were significantly associated with a worse prognosis and OS compared to KRAS-mutated patients. We also confirmed that patients diagnosed with stage IV disease, across the entire cohort, had significantly worse prognosis and survival compared to those with stage I to III disease.
Limitations. In addition to our study’s strengths, several limitations exist. This study was retrospective, hypothesis-generating, and subject to potential biases associated with RWE. The multivariate analysis focuses on prognosis and OS risk and does not include specific treatment regimens or other potential KRAS comutations that could affect survival outcomes.
Conclusion
This analysis provides valuable insights into the prognostic impact of KRAS and SMARCA4 mutations, including their comutations, on survival outcomes in metastatic NSCLC. Utilizing the AACR GENIE BPC NSCLC 2.0-public dataset, we demonstrated that KRAS/SMARCA4 class 1 comutations were significantly associated with worse OS compared to KRAS mutations alone in patients with NSCLC. These findings reinforce the importance of routine genomic profiling in patients with NSCLC to optimize precision oncology treatment strategies further.
Our results highlight the biological complexity of SMARCA4 class 1 versus class 2 mutations and their role in tumor progression when comutated with KRAS in patients with NSCLC. Future prospective studies should include KRAS/SMARCA4 class 1 and KRAS/SMARCA4 class 2 cohorts to validate the prognostic effects and potential predictive value of these comutations on survival outcomes in this high-risk NSCLC population.
Acknowledgements
A special thank you to AM for all her support. The authors would like to acknowledge the American Association for Cancer Research and its financial and material support in the development of the AACR Project GENIE registry, as well as members of the consortium for their commitment to data sharing. Interpretations are the responsibility of the study authors.
Author Contributions
Conceptualization: PM, JW, LB, and DI; Methodology: Y-BW, PM, and JW; Software: Y-BW; Formal Analysis: Y-BW; Investigation: PM; Resources: PM; Data Curation: PM; Writing—Original Draft Preparation: PM; Writing—Review and Editing: PM, JW, LB, Y-BW, and DI; Supervision: JW, LB, and DI. All authors have read and agreed to the published version of the manuscript.
Data Availability Statement
All data utilized in this analysis were obtained from the AACR Project GENIE BPC database. The dataset is publicly available through the AACR GENIE Data Commons (https://genie.cbioportal.org/). The specific dataset version employed is [Version 2.0-public]. Researchers can access the dataset following the instructions provided on the AACR GENIE website.
References
- World Health Organization Lung Cancer. Accessed 13 Nov 2024. https://www.who.int/news-room/fact-sheets/detail/lung-cancer
- Thandra K, Barsouk A, Saginala K, et al. Epidemiology of lung cancer. Contemp Oncol (Pozn)l. 2021;25(1):45–52.
- Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72(1):7–33.
- Ganti AK, Klein AB, Cotarla I, et al. Update of incidence, prevalence, survival, and initial treatment in patients with non-small cell lung cancer in the US. JAMA Oncol. 2021;7(12):
1824–1832. - Surveillance Research Program, National Cancer Institute SEER*Explorer: an interactive website for SEER cancer statistics. Accessed 23 Nov 2024. https://seer.cancer.gov/statistics-network/explorer/
- Soria JC, Ohe Y, Vansteenkiste J, et al. Osimertinib in untreated EGFR-mutated advanced non-small-cell lung cancer. N Engl J Med. 2018;378(2):113–125.
- Jänne PA, Riely GJ, Gadgeel SM, et al. Adagrasib in non-small-cell lung cancer harboring a KRASG12C mutation. N Engl J Med. 2022;387(2):120–131.
- Jordan EJ, Kim HR, Arcila ME, et al. Prospective comprehensive molecular characterization of lung adenocarcinomas for efficient patient matching to approved and emerging therapies. Cancer Discov. 2017;7(6):596–609.
- Pan W, Yang Y, Zhu H, et al. KRAS mutation is a weak, but valid predictor for poor prognosis and treatment outcomes in NSCLC: a meta-analysis of 41 studies. Oncotarget. 2016;7(7):8373–8388.
- Manolakos P, Ward LD. A critical review of the prognostic and predictive implications of KRAS and STK11 mutations and co-mutations in metastatic non-small lung cancer. J Pers Med. 2023;13(6):1010.
- Skoulidis F, Goldberg ME, Greenawalt DM, et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov. 2018;8(7):822–835.
- Dagogo-Jack I, Schrock AB, Kem M, et al. Clinicopathologic characteristics of BRG1-deficient NSCLC. J Thorac Oncol. 2020;15(5):766–776.
- Schoenfeld AJ, Bandlamudi C, Lavery JA, et al. The genomic landscape of SMARCA4 alterations and associations with outcomes in patients with lung cancer. Clin Cancer Res. 2020;26(21):5701–5708.
- Herzberg B, Gandhi N, Henick B, et al. Effects of mutations in SWI/SNF subunits on context-specific prognosis in driver positive and driver negative NSCLC. Proceedings of the 2023 American Society of Clinical Oncology Annual Meeting; Chicago, IL, US; 2–6 Jun 2023; Abstract 9039.
- Liang X, Gao X, Wang F, et al. Clinical characteristics and prognostic analysis of SMARCA4-deficient non-small cell lung cancer. Cancer Med. 2023;12(13):14171–14182.
- Manolakos P, Boccuto L, Ivankovic DS. A critical review of the impact of SMARCA4 mutations on survival outcomes in non-small cell lung cancer. J Pers Med. 2024;14(7):684.
- Hodges H, Stanton B, Cermakova K, et al. Dominant-negative SMARCA4 mutants alter the accessibility landscape of tissue-unrestricted enhancers. Nat Struct Mol Biol. 2018;25(1):61–72.
- Mardinian K, Adashek JJ, Botta GP, et al. SMARCA4: implications of an altered chromatin-remodeling gene for cancer development and therapy. Mol Cancer Ther. 2021;20(12):2341–2351.
- Fernando TM, Piskol R, Bainer R, et al. Functional characterization of SMARCA4 variants identified by targeted exome-sequencing of 131,668 cancer patients. Nat Commun. 2020;11(1):5551.
- de Bruijn I, Kundra R, Mastrogiacomo B, et al. Analysis and visualization of longitudinal genomic and clinical data from the AACR Project GENIE Biopharma Collaborative in cBioPortal. Cancer Res. 2023;83(23):3861–3867.
- Alessi JV, Ricciuti B, Spurr LF, et al. SMARCA4 and other SWItch/Sucrose Non-fermentable family genomic alterations in NSCLC: clinicopathologic characteristics and outcomes to immune checkpoint inhibition. J Thorac Oncol. 2021;16(7):1176–1187.
- Alessi JV, Elkrief A, Ricciuti B, et al. Clinicopathologic and genomic factors impacting efficacy of first-line chemoimmunotherapy in advanced NSCLC. J Thorac Oncol. 2023;18(6):731–743.
- Negrao MV, Araujo HA, Lamberti G, et al. Comutations and KRASG12C inhibitor efficacy in advanced NSCLC. Cancer Discov. 2023;13(7):
1556–1571. - Boiarsky D, Lydon CA, Chambers ES, et al. Molecular markers of metastatic disease in KRAS-mutant lung adenocarcinoma. Ann Oncol. 2023;34(7):589–604.
- Liu L, Ahmed T, Petty WJ, et al. SMARCA4 mutations in KRAS-mutant lung adenocarcinoma: a multi-cohort analysis. Mol Oncol. 2021;15(2):462–472.
- Choudhury NJ, Lavery JA, Brown S, et al. The GENIE BPC NSCLC cohort: a real-world repository integrating standardized clinical and genomic data for 1,846 patients with non-small cell lung cancer. Clin Cancer Res. 2023;29(17):3418–3428.
- AACR Project GENIE Consortium AACR Project GENIE: powering precision medicine through an international consortium. Cancer Discov. 2017;7(8):818–831.
- Lavery JA, Lepisto EM, Brown S, et al. A scalable quality assurance process for curating oncology electronic health records: The Project GENIE Biopharma Collaborative approach. JCO Clin Cancer Inform. 2022;6:e2100105.
- Wankhede D, Grover S, Hofman P. SMARCA4 alterations in non-small cell lung cancer: a systematic review and meta-analysis. J Clin Pathol. 2024;77(7):457–463.
- Alver BH, Kim KH, Lu P, et al. The SWI/SNF chromatin remodeling complex is required for maintenance of lineage specific enhancers. Nat Commun. 2017;8:14648.
- Ito K, Hulse M, Agarwal A, et al. Discovery of PRT3789, a selective SMARCA2 degrader. Proceedings of the AACR-NCI-EORTC International Conference on Molecular Targets and Cancer Therapeutics, Hynes Convention Center; Boston, MA, US; 11–15 Oct 2023.
- Dagogo-Jack I, Dowlati A, Guo R, et al. A Phase 1 study of PRT3789, a potent and selective degrader of SMARCA2 in patients with advanced or metastatic solid tumors and a SMARCA4 Mutation. Proceedings of the Annual European Society for Medical Oncology Congress (ESMO); Madrid, Spain; 20–24 Oct 2023.
- Yap TA, Tolcher AW, Plummer R, et al. First-in-human study of the ataxia telangiectasia and Rad3-related (ATR) inhibitor tuvusertib (M1774) as monotherapy in patients with solid tumors. Clin Cancer Res. 2024;30(10):2057–2067.
- Paz-Ares L, Cappuzzo F, Yamamoto N, et al. 104TiP—Phase Ib/IIa study of ATR inhibitor tuvusertib + anti-PD-1 cemiplimab in patients with advanced non-squamous non-small cell lung cancer (NSCLC) that has progressed on prior anti-PD-(L)1 and platinum-based therapies. Ann Oncol. 2024;9(Suppl 3):102683.